|
9/22/2023 0 Comments Bcftools filter vcf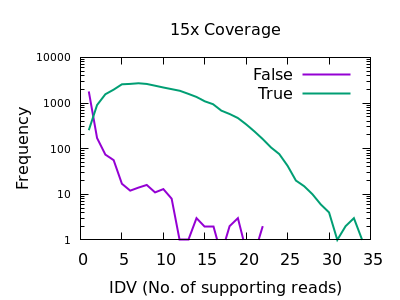
The main commands to extract the specific SNP/INDEL markers are as the folowings:īcftools query -H -i'%TYPE="indels"' -f "%CHROM %POS %REF %ALT %DP %QUAL %AF %AC %AN %TYPEīcftools query -H -i'%TYPE="snps"' -f "%CHROM %POS %REF %ALT %DP %QUAL %AF %AC %AN %TYPEĢ. Call bcftool to extract the specific SNP/INDEL Markers GMEF mainly include the following functional sub-procedures.ġ. 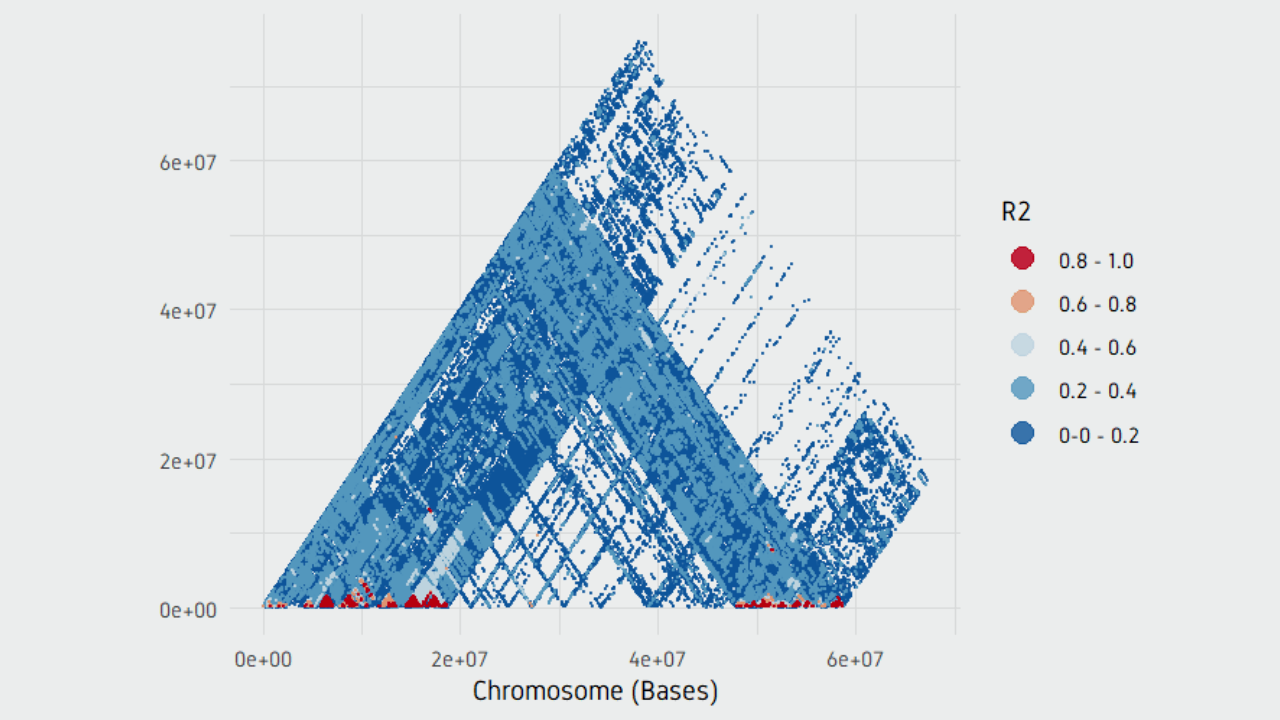
The first 3 criterias are calculated as a percentages. The specific criterias include: MAF(Minum Allele Frequencey), Heterogenous Genotype Ratio, Unknown Genotype Ratio, and the Allele Length Difference. The user can specific the marker type as SNP or/and INDEL, whether need to exclude the markers from chloroplast or/and scaffolds, whether the markers meet the specific criterias. The users only need to select the required variant call file, configure the parameters and submit. In GMEF, we have hosted several genetic variant data including the medicago hapmap data. GMEF is a web pipeline, which is aiming to extract and filter the genotype markers from the original genetic variant file, such as.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
